Postdocs
Németh Tibor
Project title : Creating an overexpression library and developing an inducible gene expression system in Candida parapsilosis
To study the interaction between mammalian macrophages and Candida parapsilosis an in vitro model system was set up and optimised involving J774.2 murine cells. We established that these phagocytes were capable of uptake and eliminate the yeast cells. To characterise transcriptional changes of the host upon interaction microarray analysis and quantitative real-time PCR were performed. The C. parapsilosis – host interaction was examined by using THP-1 human monocytes as well. Up- and downregualted genes of both the host and the pathogen were identified by RNAseq analysis at four different time-points. Our major goal is to create a BAR-coded overexpression library in C. parpasilosis based on these data to study virulence in vitro and in vivo. To create mutants in such a large scale LifeTechnolgies Gateway system is applied. This approach is also used for creating reintegrant mutants for C. parapsilosis knock-out library and to adopt TETON-TETOFF system optimised for C. albicans. This approach enables us to regulate the expression (overexpress or silence) of any genes under in vitro or in vivo circumstances at any timeponts of the interaction.
Metal ions are essential nutrients for both the host and the fungus. The way how pathogens can obtain specific ions upon infection, when accessible metal ions are limited, is still not known in every details. Previous data revealed that C. albicans possessed a “siderophore-like” scavenger-transporter system that was responsible for uptake zinc ions upon in vitro infection. This so-called “zincophore system” consists of two proteins in C. albicans, however C. parapsilosis lacks the scavenger, suggesting a different mechanism for zinc-uptake in this species. Based on homology analysis a total of six potential zinc-transporters were identified in C. parapsilosis. Our aim is to knock these genes out to reveal how this pathogen is capable of acquiring zinc upon interaction with the host.
Tóth Renáta
Project title : Identification of novel virulence affecting genes via the generation of a Candida parapsilosis deletion library
Candida parapsilosis is an opportunistic pathogenic fungi species which has a constitutively diploid genome and lacks sexual cycle. In order to identify genes that are potentially involved in the fungi’s virulence, two approaches are accepted and used frequently: generation of null mutants or over-expression strains of a desired gene of interest. A part of our laboratory is currently using a previously described, auxotrophy complementation based gene knockout strategy to study gene function in Candida parapsilosis. In our recent work several fungal transcriptional factors and kinases have been identified using RNA-Seq data that were over-expressed during host-pathogen interactions. Our aim is to generate a knockout collection of these genes in order to identify new genes contributing to virulence. Besides these identified but not yet characterized factors, further ORFs have been selected and added to our ever-expanding knock out library, that might affect the fungi’s virulence.
Photo made by Tamás Petkovits and Renáta Tóth
PhD students
Csonka Katalin
Project title : Recognition of Candida parapsilosis
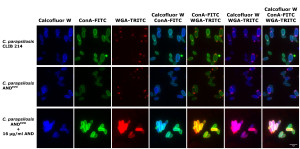
The cell wall is the first site of interaction between the fungal pathogens and host cells, influencing the immune recognition of fungal cells. The fungal cell wall constituents are pathogen-associated molecular patterns (PAMPs) that are recognized by pattern recognition receptors (PRRs) on the surface of host immune cells, for example monocytes and neutrophils in the circulation and macrophages in infected tissues.
Of the four major families of PRRs, the Toll-like receptor (TLR) family is the best studied, with TLR2 and TLR4, among others, being involved in the fungal recognition. However, antifungal immunity appears to be mediated primarily by members of the C-type lectin receptor (CLR) family, including Dectin-1 and Dectin-2. Nevertheless, the exact role of these PRRs with focus on the immunity against C. parapsilosis needs to be determined.
With using mutant cells lacking enzymes with key roles during the assembly of fungal surface components we are able to understand these cellular processes and the importance of these cell wall elements in the host-phatogen interactions.
As a PhD student, I have been working on the characterization of recognition of C. parapsilosis using different infection models such as cell culture, interaction with primary immune cells and in vivo imaging.
Dhirendra K Singh
Project title : Investigation of the innate and adaptive immune response induced by opportunistic pathogen Candida parapsilosis
My main focus of research is to identify the role of C. parapsilosis secreted aspartic acid proteases (SAPPs) in complement component-C3, C5 FH, FI and complement receptor-CR1(CD35), CR2, CR3(Cd11b/CD18 or Mac1), CR4 (CD11c/CD18) regulation. I’m also interested in identifying complement receptor-mediated signalling induced by the Sapps in macrophages and neutrophils.
Papp Csaba
Project title : Resistance mechanisms and their impact in Candida parapsilosis.
Fungal infections are emerging world wide. The leader causes of these diseases are Candida species. Treatment against Candida species are performed using azoles, echinocandins and amphotericin B. For these microorganisms it is essential to evolve resistance against the most commonly used antimycotics. However, many studies have showed the fitness and virulence cost of drug resistant Candida strains. My work is focusing on the mechanisms of resistance development and their influence on virulence and fitness.
Tanmoy Chakraborty
Project title : The role of prostaglandin-like molecules in host-pathogen interaction of Candida parapsilosis
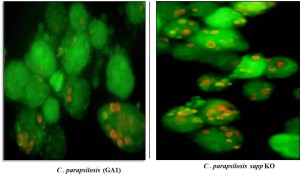
Candida parapsilosis is one of the major human fungal pathogen, which primarily causes invasive infection among low birth weight infants. Despite its increasing prevalence very little is known about the mechanisms involved in its interaction with the human host. The eicosanoids are 20C long chain fatty acid molecules which are mainly derived from arachidonic acid. They are the oxygenated metabolites of arachidonic acid produced mostly by the enzymatic action of COX and LOX enzyme. There are different eicosanoids produced in the mammalian host belong to the different subfamilies, which include prostaglandins, thromboxane, leukotrienes, lipoxins, resolvins and HETEs (Hydroperooxy-eicosatraenoic acid). These eicosanoids are the bioactive signaling lipid molecules, which plays a major role not only in infection and inflammation but also in resolution of inflammation. Although fungi lack the homologue of Cox/Lox enzymes still they can produce prostaglandins from exogenous arachidonic acid. In recent years it has been shown by different groups that human pathogenic fungi like Candida albicans, Candida parasilosis, Cryptococcus neoformans, Paracoccidioides brasiliensis and Aspergillus fumigatus also produce prostaglandin molecules which is similar to the host. In case of C. albicans, it has been found that two genes namely OLE2 (Fatty acid desaturase homologue) and FET3 (Multicopper oxidase homologue) is involved the prostaglandinE2 (PGE2) production. However, it has been shown from our lab that, unlike the closely related pathogenic fungus C. albicans, OLE2 (fatty acid desaturase gene) is not involved in the biosynthesis of Prostaglandin in C. parapsilosis. The current work involves the analysis of the novel genes or pathways for different eicosanoids production in C. parapsilosis including prostaglandins and how these fungal lipid mediators are involved in inflammation and host pathogen interaction during candidiasis.
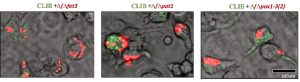
To identify genes for eicosanoid biosynthesis in C. parapsilosis, we performed RNA-seq analysis after growing the cells in presence of arachidonic acid, the main precursor for eicosanoid production. The sequence analysis (in collaboration with Dr. Toni Gabaldon’s Lab, CRG, Barcelona) showed that total 68 genes were upregulated and 83 genes were downregulated. From the upregulated genes we found that 14.5% genes are involved in lipid metabolism. Six probable candidate genes were selected by GO term analyses which are involved in the lipid metabolism process. Knock out (KO) mutants for those 6 genes were generated by fusion PCR technique. The 6 mutants were then analyzed for the eicosanoid production. In the beginning mainly the cell free supernatant was analyzed after growing the cells in presence of 100uM arachidonic acid. The eicosanoid analysis was done in the LC/MS facility in collaboration with Dr. Martin Giera’s lab, LUMC, Leiden. The analysis showed that other than different prostaglandins (PGD2, PGE2 and 15-K-PGE2) C. parapsilosis also produce lipoxinA4 molecules, which is a novel finding according to our knowledge. From the 6 KO mutants 3 of them showed significant reduction in PGE2 and LipoxinA4 production compared to the wild type (WT) strain. Currently samples were prepared to analyze the internal eicosanoid from the mutant strains.
As the lipid mediators also have effect on the morphogenesis and biofilm production of C. albicans, the mutants were analyzed for their fitness after growing them in different temperature, pH, stress source and pseudo hypha inducing media. Few of the mutants showed prominent difference in some of the growth condition. Currently the mutants are also studied to check whether they have any difference in their ability to form biofilms, as C. parapsilosis also can biofilm, one of the pathogenic trait of this organism.
Finally, to study the role of the fungal lipid mediators in inflammation and host pathogen interaction during candida infection, human primary macrophage cells are being used as a model system (PBMC-Peripheral Blood Mononuclear Cell and PBMC derived macrophages). The recent results showed that the PBMC-DMs are more efficient in killing and phagocytose the eicosanoid KO mutants compared to the WT C. parapsilosis strain. In the ongoing experiments, the mutants are compared with the WT strain for their ability to generate inflammatory response by analyzing different pro and anti-inflammatory cytokines and chemokines (IL1β, TNFα, IL10 and IL8) by ELISA after infecting both PBMC and PBMC-DM. Preliminary results showed that the KO mutants produce higher amount of anti-inflammatory cytokine IL10 which correlates the lower production of pro-inflammatory cytokine TNFα production. The future plan is to analyze the mutants for their virulence using mouse model of systemic infection.
Zajta Erik
Project title: Syk/CARD9 signaling in Candida parapsilosis infection
This project focuses on Syk and CARD9. These signaling components have important functions in multiple innate immunological processes. Recognition of some well studied fungal pathogens by the innate immune system is heavily dependent on Syk/CARD9 mediated sinaling. This is initiated by C-type lectin receptors Dectin-1, Dectin-2 and Mincle which bind fungal cell wall components such as β-glucans and N-linked mannans.
Within this project, my aim is to study the role of Syk mediated signaling during Candida parapsilosis infection.

Máté Vadovics
Project title: The microbiome and cancer
A large number of commensal microbial species reside in the human body that have co-evolved with the human genome and adopted to the host immune system. Previously it has been shown that defects in regulatory processes or alterations in the composition of microbiome can lead to various diseases, including cancer. Even though the human bacteriome is well-investigated, our knowledge is limited about the composition and function of the residential fungal microflora or mycobiome.
The most frequent opportunistic fungal infection in the oral cavity is candidiasis. Previous studies have shown that the number of Candida cells in this particular niche is significantly higher in patients with oral squamous cell carcinoma (OSCC) compared to healthy individuals. It was suggested that tumour-induced immunosuppression could cause fungal overgrowth and contribute to the development of an infection. However, the etiological relationship between oral cancer and Candida infections is still a matter of debate. Our aim is to examine the influence of fungal or bacterial infections on the metastatic activity of cancerous cells, furthermore the effects of extracellular vesicles on host cells released by Candida albicans and Candida parapsilosis.
Sára Pál
Project title: Examination of virulence-related genes in the Candida parapsilosis with the help of overexpression and knock out mutant collections.
The aim of our project is to create an overexpression (OE) mutant collection in order to investigate the role of fungal ORFs potentially involved in C. parapsilosis virulence. Further in vivo and in vitro examinations are performed. We would like to compare these new datas with former results derive from the investigation of our knock out collection made by Renáta Tóth to gain new information about the pathogenicity of this model organism. We use the so called Gateway cloning technology (Thermo Fisher Scientific) to create OE strains supervising by Tibor Németh.
In parallel we would like to adapt some already existing CRISPR/Cas9 constructs in our laboratory to create new mutant strains also under Tibor Németh’s leadership.
MSc students
Benedikta Balla
Supervisor: Csaba G Papp
Regina E Bernátsky
Supervisor: Tibor Németh
Mónika Homolya
Supervisor: Renáta Tóth
Project title : Analysis of a Candida parapsilosis hyper-filamentous mutant strain
BSc students
Katica Kocsis
Supervisor: Renáta Tóth
Project title : Characterization of Candida parapsilosis cell wall over-expression mutant strains
Flóra F Bohner
Supervisor: Renáta Tóth
Project title : General characterization of Candida parapsilosis deletion mutant strains
Gergő Drabbant
Supervisor: Renáta Tóth
Project title : Phenotypic analysis of Candida parapsilosis transcriptional factor over-expressing mutant strains
Tamás Takács
Supervisor: Tibor Németh
Project title : Phenotypic characterisation of zinc transporters in Candida parapsilosis
Máté Nyúl
Supervisor: Erik Zajta